The following in a data model for an animal classification system. This system contains multimedia records for many types of animal. The system allows for the construction of a variety of classifications by using Node objects. By adding new classifications or exploring existing ones, users can examine a variety of relationships between animals. They can using alternate classifications, or even intersect different classifications, depending on their current information need. Possible users include junior high students, amateur enthusiasts, researchers, and zoological taxonomists.
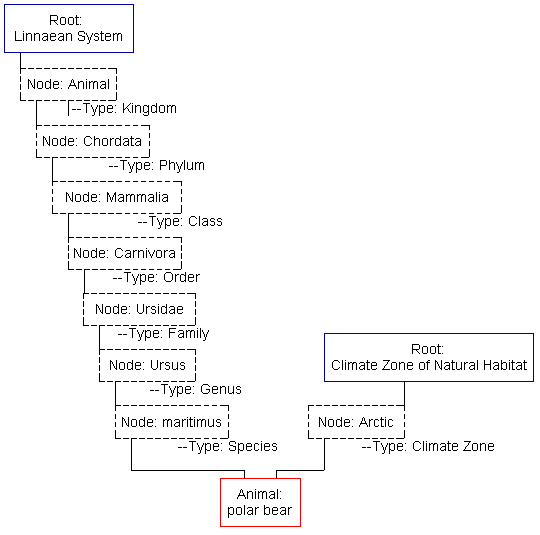
Because of the redundant/recursive use of nodes, this system can be a rather hard to imagine. Here's an illustrative example:

As you can see, the root-node architecture allows for the construction of practically any number of classification systems. There can be any number of node levels; indeed, the Climate Zone classification only uses one such level. (Other likely nodes may be "desert", "rain forest", "tundra", "marine", etc.) Also, I have only shown one branch here of a tree-like structure. At each level of the Linnaean System there are a number of other nodes--many Families under the same Order, many Species under the same Genus, etc. These systems all use the same animal records. (Not every classification system will use all the animal records, however; for example, perhaps someone wishes only to classify snakes on the basis of their venom and how they interact with known anti-venoms.)
Ideally, queries will be able to explore multiple classifications and combine the results. For example, imagine someone compiling an identification guide for marine mammals. Using only the Linnaean System, they would have to sift through all mammals in order to find groups such as whales and dolphins (Order="Cetacea"). Using only a Climate Zone classification that includes a "Marine" zone, they would be swamped with fish, sponge, coral, and other irrelevant animal records. However, if they could intersect these results, they would find whales and dolphins, as well as sea otters, dugongs, seals, and other marine mammals not easily found traversing only one classification system or the other.
Here are some other possible information needs that might be served by this system:
I hope these examples shed some light on the nature of this system.
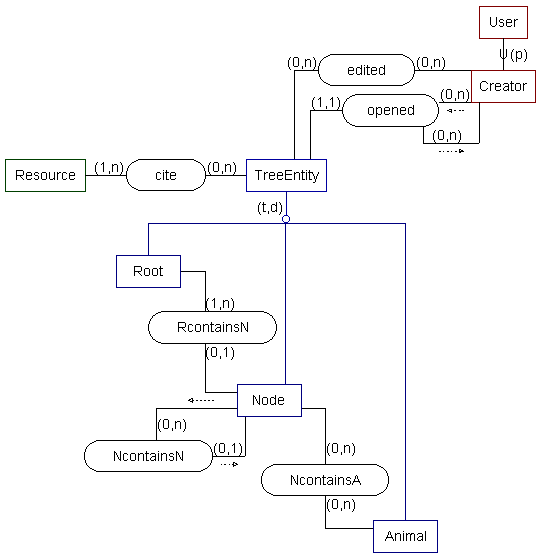
-- SSM model --
| Animal | |
|---|---|
| species_name | <vchar(60)> -- the Latin Genus-species name for this animal |
| common_name | <vchar(60)> -- the common English name for this animal. |
| keywords | <vchar(200)> -- keywords describing this animal to aid in searching |
| Description | <text> |
| Image | filename: <vchar(30)> -- the name of the image file pixel-width: <dec(4)> -- the width of the image in pixels pixel-height: <dec(4)> -- the height of the image in pixels format: <vchar(4)> -- {JPEG | GIF | PNG | BMP} comments: <vchar(500)> -- any comments on the image image: <image> |
The Animal record are the foundation of the system. Each record is like an encyclopedia entry, with photo, text description, and resources for further information.
I have learned an animal's common name may be misleading depending on which field guide one uses or on one's nationality. Species names are, by design, original, unique, and universal (as much as possible, anyway). I have included the species name in the animal record because I think it will probably be used often for direct retrieval.
The complex Image object includes metadata specific to the format of the image. This will be useful for generating HTML on-the-fly. Comments could be sent as ALT text.
| Root | |
|---|---|
| id | <dec(12)> -- a unique system id for this entity |
| classif_name | <vchar(60)> -- the name of the classification system stemming from this root |
| keywords | <vchar(200)> -- keywords describing this classification to aid in searching |
| scope | <char(3)> {ALL | SUB} -- whether this classification aims to include all animals, or just a subset |
| description | <vchar(1000)> -- a description of this classification system |
Root serves to placehold the top of a classification system. For example, the well-known Kingdom-Phylum-Order-Family-Genus-Species classification is known as the Linnaean System. If this system was being modeled, Root.classif_name would likely equal "Linnaean System".
| Node | |
|---|---|
| id | <dec(12)> -- a unique system id for this entity |
| taxon_name | <vchar(60)> -- the name of the particular taxon or group or classification level represented by this node |
| type | <vchar(40)> -- describes the type of taxon or the level within the classification system |
| keywords | <vchar(200)> -- keywords describing this node to aid in searching |
| Description | <vchar(1000)> -- a description of this entity |
The nodes will form the bulk of the classification structure and possibly be the most common record type in the database. Type serves to describe the level or taxon of the node. For example, if this were a node somewhere between a "Linnaean System" Root and a "polar bear" Animal record, the taxon_name may equal "Ursus" and the type equal "Genus." (It would then contain another node, taxon_name="maritimus" and type="Species", which would then contain the "polar bear" record.)
Keywords can contain common synonyms for taxon_names. For example, the Genus Ursus node should certainly contain the keyword "bears".
| TreeEntity | |
|---|---|
| id | <dec(12)> -- a unique system id for this entity |
TreeEntity is simply an abstract superclass of Animal, Node, and Root. It greatly simplifies the SSM model by concentrating all the common relationships with Resources and Creators.
| Resource | |
|---|---|
| id | <dec(12)> |
| author | <vchar(50)> |
| title | <vchar(200)> |
| publisher | <vchar(60)> |
| location_or_URL | <vchar(100)> |
| pub_date | <date> |
| notes | <vchar(250)> -- notes or comments about this resource |
I think this is pretty clear. It's just your basic, simple bibliographic record.
| User | |
|---|---|
| username | <vchar(20)> -- a unique username |
| age | <dec(2)> -- assumes that centenarians are not web-surfing taxonomists |
| education | {ELEMENTARY | MIDDLE | HIGH SCHOOL | COLLEGE | GRAD | TEACHER | PROFESSOR} |
| nationality | <vchar(30)> |
| <vchar(30)> | |
| created | <date> |
| last_accessed | <date> |
| time_inactive | <derived> -- how long it has been since this user logged in |
These records help accumulate information about who is using the system. Note that there is a username but no password. Users can merely search the system. Since this is not a security issue, there seems no need for passwords that people will just forget anyway.
| Creator | |
|---|---|
| password | <vchar(12)> |
| name | <vchar(50)> |
| address | <vchar(120)> |
| release_prefs | {USERNAME | NAME | EMAIL | ADDRESS} -- how much information to release to other users; each choice includes those before it. |
| greeting | <vchar(750)> -- an introductory message to other users of the system |
A Creator is a User who can develop new classification schemes or add animal records. Since the system keeps track of their work, they need a password to login.
| RcontainsN | |
|---|---|
| E1: Root | (1,n) -- A Root must contain some Nodes. |
| E2: Node | (0,1) -- A Node may or may not be contained by a Root |
| NcontainsN | |
|---|---|
| E1: Node | (0,n) -- A Node may or may not contain other Nodes. |
| E2: Node | (0,1) -- A Node may be contained by another Node. |
| NcontainsA | |
|---|---|
| E1: Node | (0,n) -- A Node may or may not contain Animals |
| E2: Animal | (0,n) -- An Animal may be contained by any number of Nodes, or none at all. (Eventually, all Animals will be contained by Nodes.) |
| cite | |
|---|---|
| E1: TreeEntity | (0,n) -- An entity may cite any number of Resources. |
| E2: Resource | (1,n) -- A Resource must be cited at least once. |
| edited | |
|---|---|
| E1: Creator | (0,n) -- A Creator may edit any number of entities. |
| E2: TreeEntity | (0,n) -- An entity can be edited any number of times. |
| time | <time> -- The time (including date) the new TreeEntity was edited. |
| opened | |
|---|---|
| E1: Creator | (0,n) -- A creator may not have created an entity yet; if he has, he can create any number of them |
| E2: TreeEntity | (1,1) -- Each set of access permissions is related to only a single Entity. |
| E3: Creators | (0,n) -- A Creator grants access to any number of Creators, 0 being all. |
| time | <time> -- The time (including date) the new TreeEntity was added. |
Since all classifications and animal records are input by Creators, there seems a need for recognition for and control of these creations. The opened relationship is a combination of a creates and a grantsAccess relationship. First of all, the Creator has a link to the entity she has added that identifies her as the original creator. Then, because of this relationship, she also has the power to grant access to other Creators. Those Creators with such granted access to an Entity are then able to edit that Entity. Hopefully Creators will grant access to all other Creators. However, they can also restrict access to only themselves, or any list in between.
Granting access to 0 Creators means granting it to all. This seemed the easiest way to include an "Open to All" option without trying to maintain an index of all possible Creators.
Comments or questions are appreciated!
Thank you for reading this far! The following is a list of notes, comments and thoughts concerning current problems and future developments. Though not officially part of this report, comments on these points would be most helpful.
Animal:
Root:
Resource:
User:
contains:
opened:
If this system was ever implemented, it would be v1.0. Once it was patched and debugged and working at a basic level, it would be time to start thinking about enhancements and extra features. Here are some of those possible features:
| ~ztomasze Index:
LIS: Initial Data Model http://www2.hawaii.edu/~ztomasze |
Last Edited: 27 Oct 2001 ©2001 by Z. Tomaszewski. |